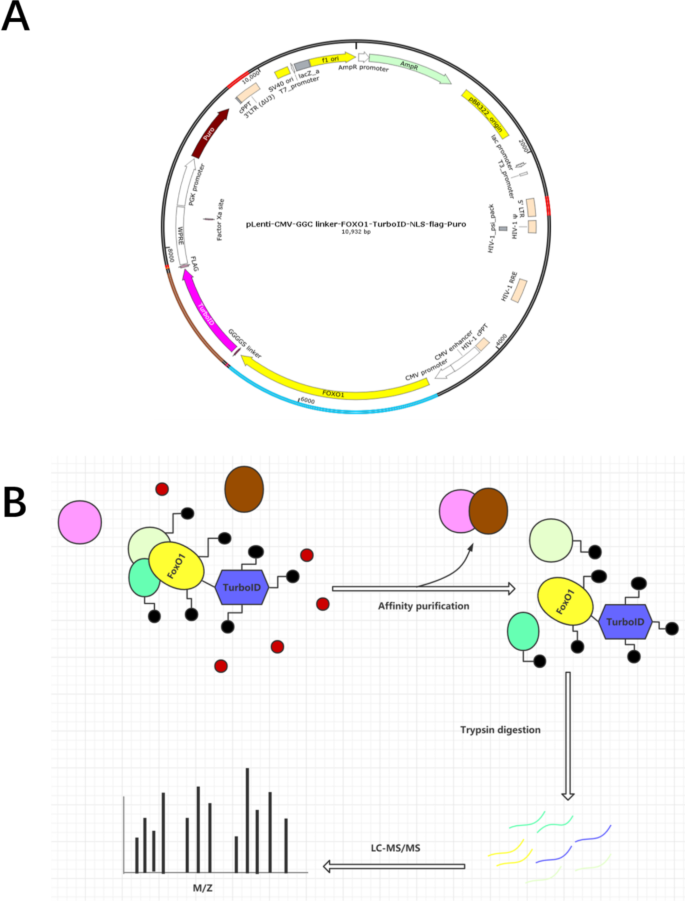
Study of FOXO1-interacting proteins using TurboID-based proximity labeling technology | BMC Genomics | Full Text

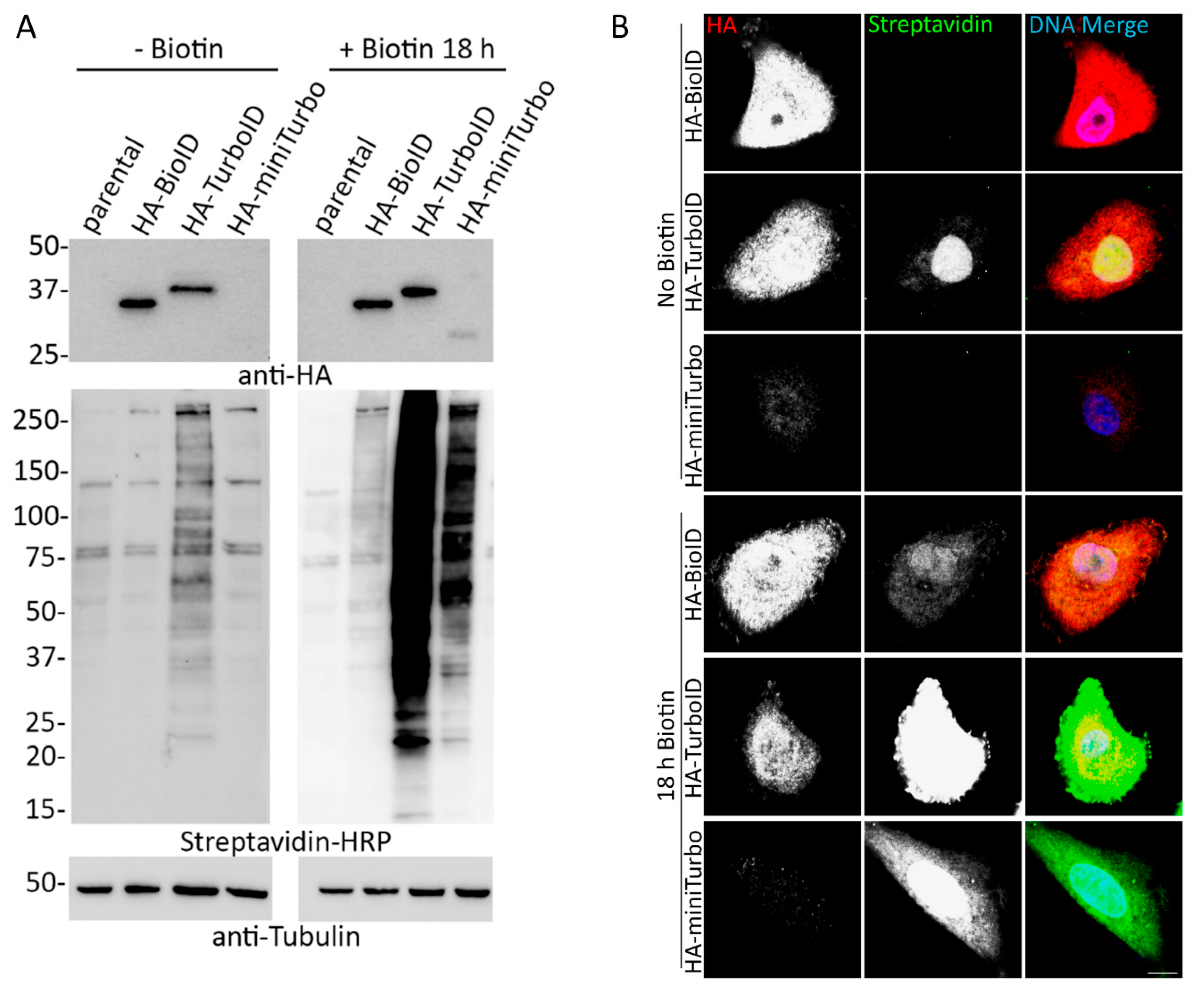
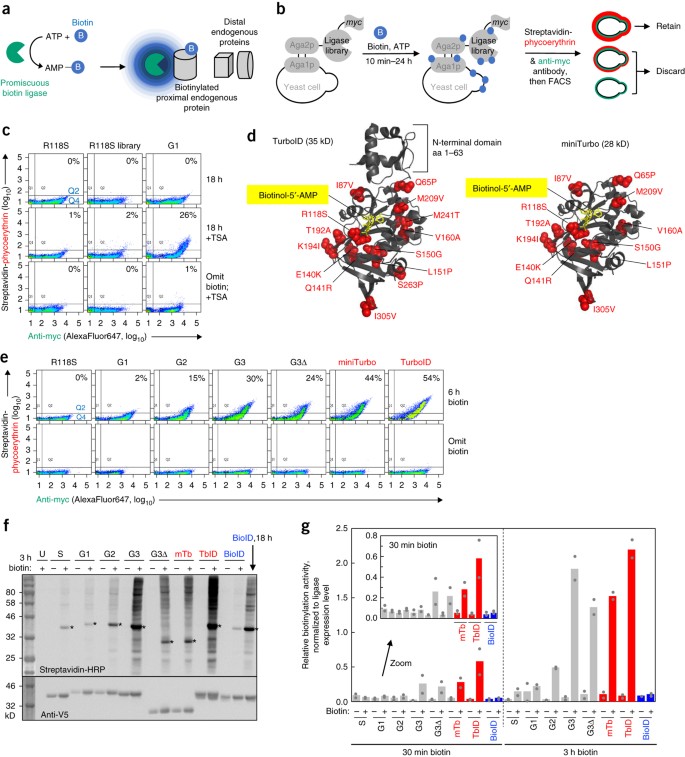
Directed evolution of TurboID for efficient proximity labeling in living cells and organisms | bioRxiv

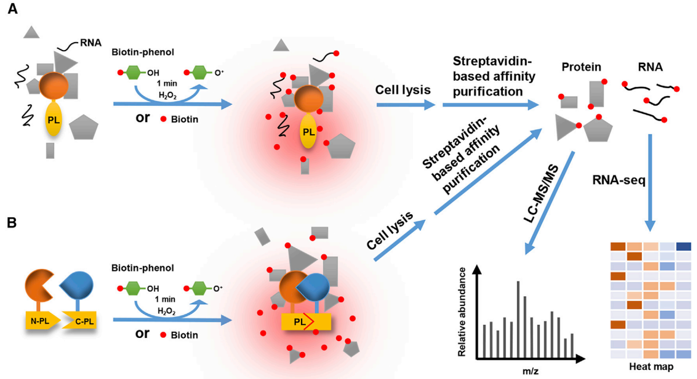
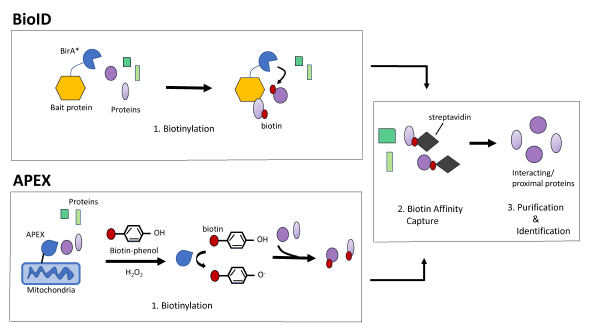
figure supplement 1. Protein proximity labeling by biotin ligases BioID... | Download Scientific Diagram
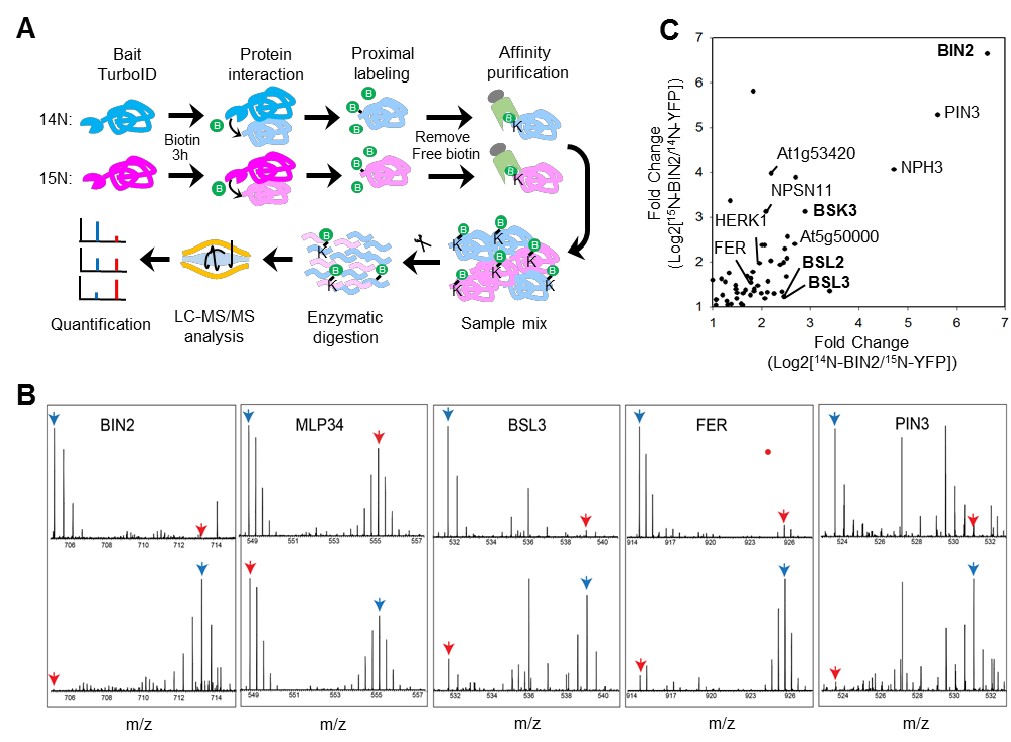
Plantae | Application of TurboID-mediated proximity labeling for mapping a GSK3 Kinase signaling network in Arabidopsis (bioRxiv) | Plantae

Thiol-Cleavable Biotin for Chemical and Enzymatic Biotinylation and Its Application to Mitochondrial TurboID Proteomics | Journal of the American Society for Mass Spectrometry

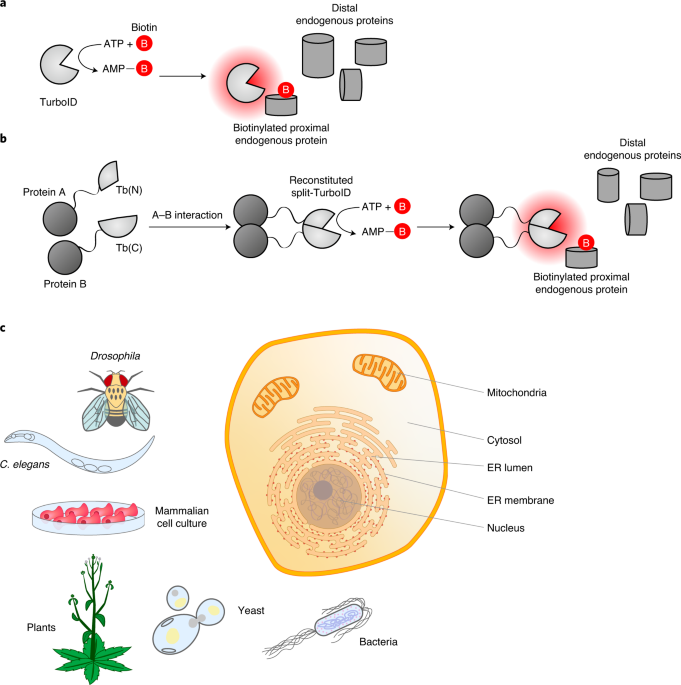
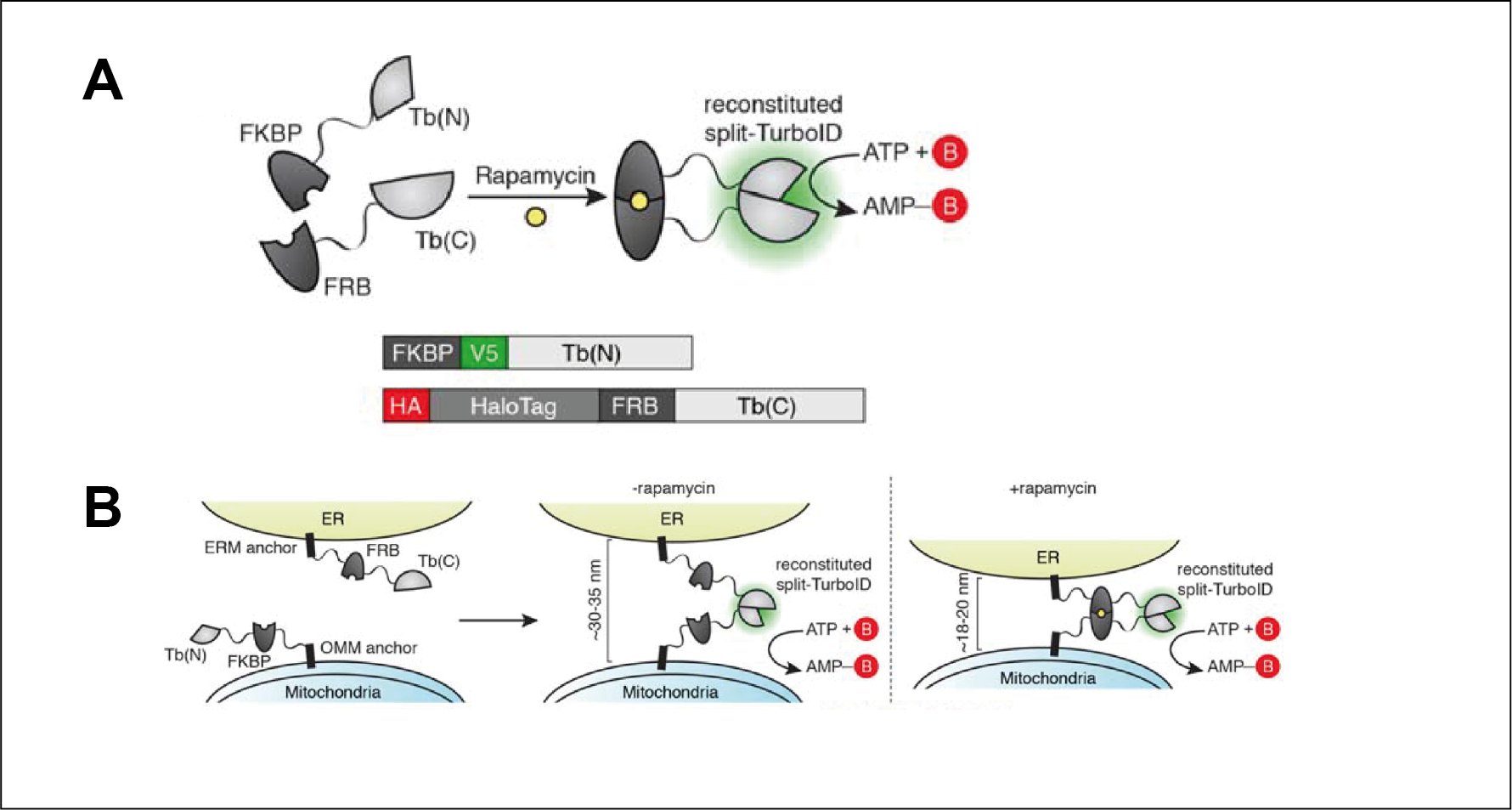
TurboID and miniTurbo- engineered promiscuous biotin ligases for efficient proximity labeling in living cells and organisms | Explore Technologies

Thiol-Cleavable Biotin for Chemical and Enzymatic Biotinylation and Its Application to Mitochondrial TurboID Proteomics | Journal of the American Society for Mass Spectrometry

Establishment of in vivo proximity labeling with biotin using TurboID in the filamentous fungus Sordaria macrospora | Scientific Reports
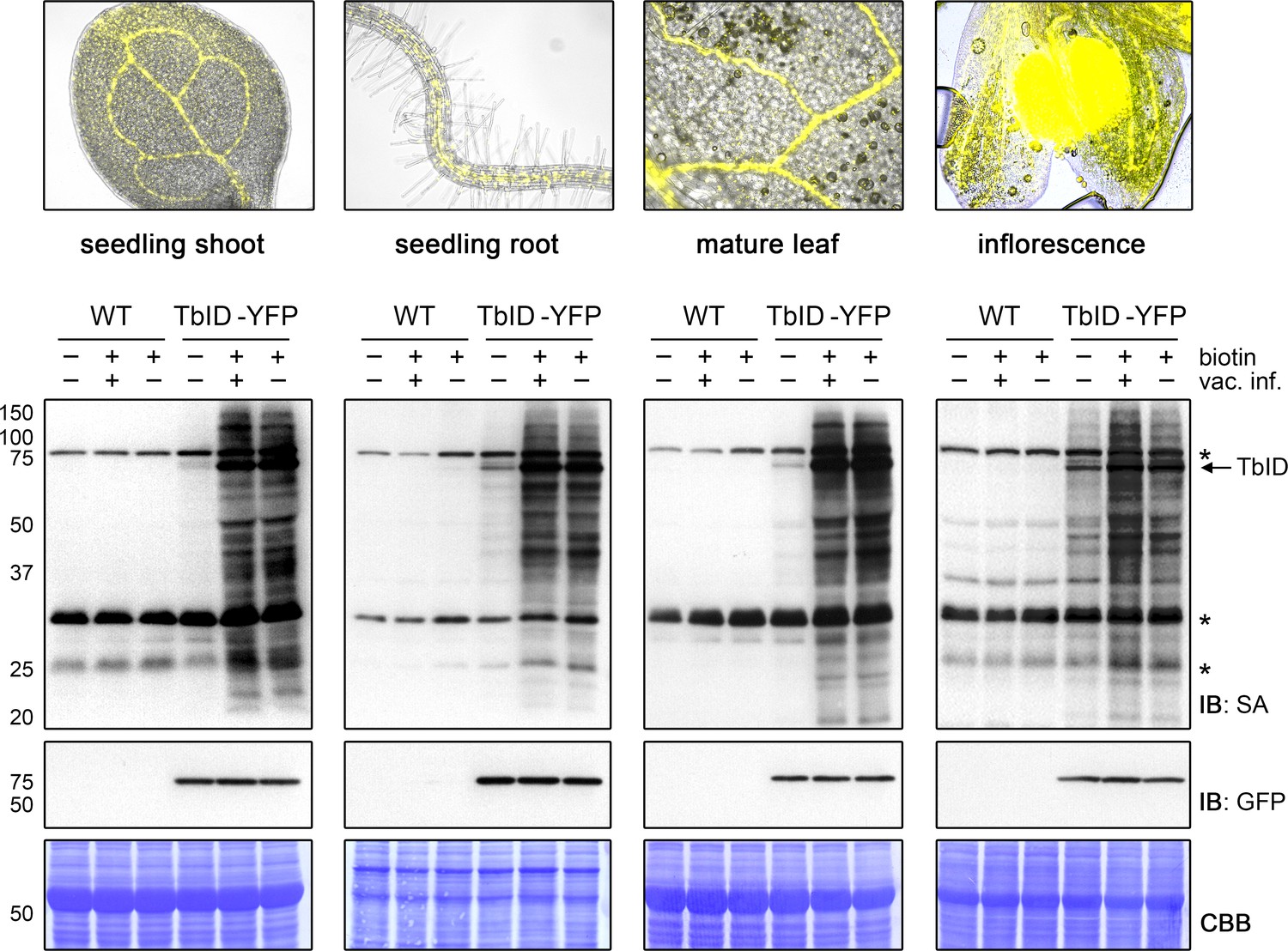
Proximity labeling of protein complexes and cell-type-specific organellar proteomes in Arabidopsis enabled by TurboID | eLife

TurboID-based proximity labeling accelerates discovery of neighboring proteins in plants: Trends in Plant Science