Proximity RNA labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules | bioRxiv

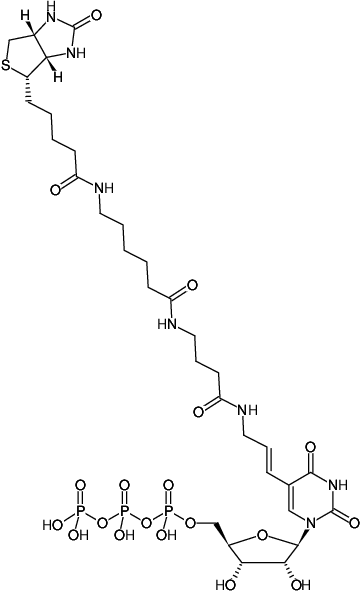
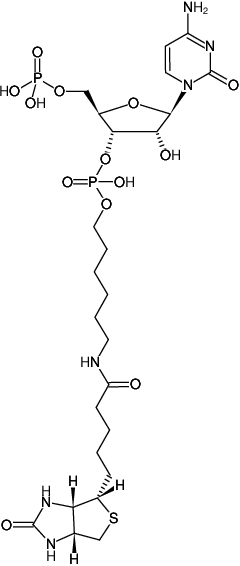
Synthesis of biotin–AMP conjugate for 5′ biotin labeling of RNA through one-step in vitro transcription | Nature Protocols
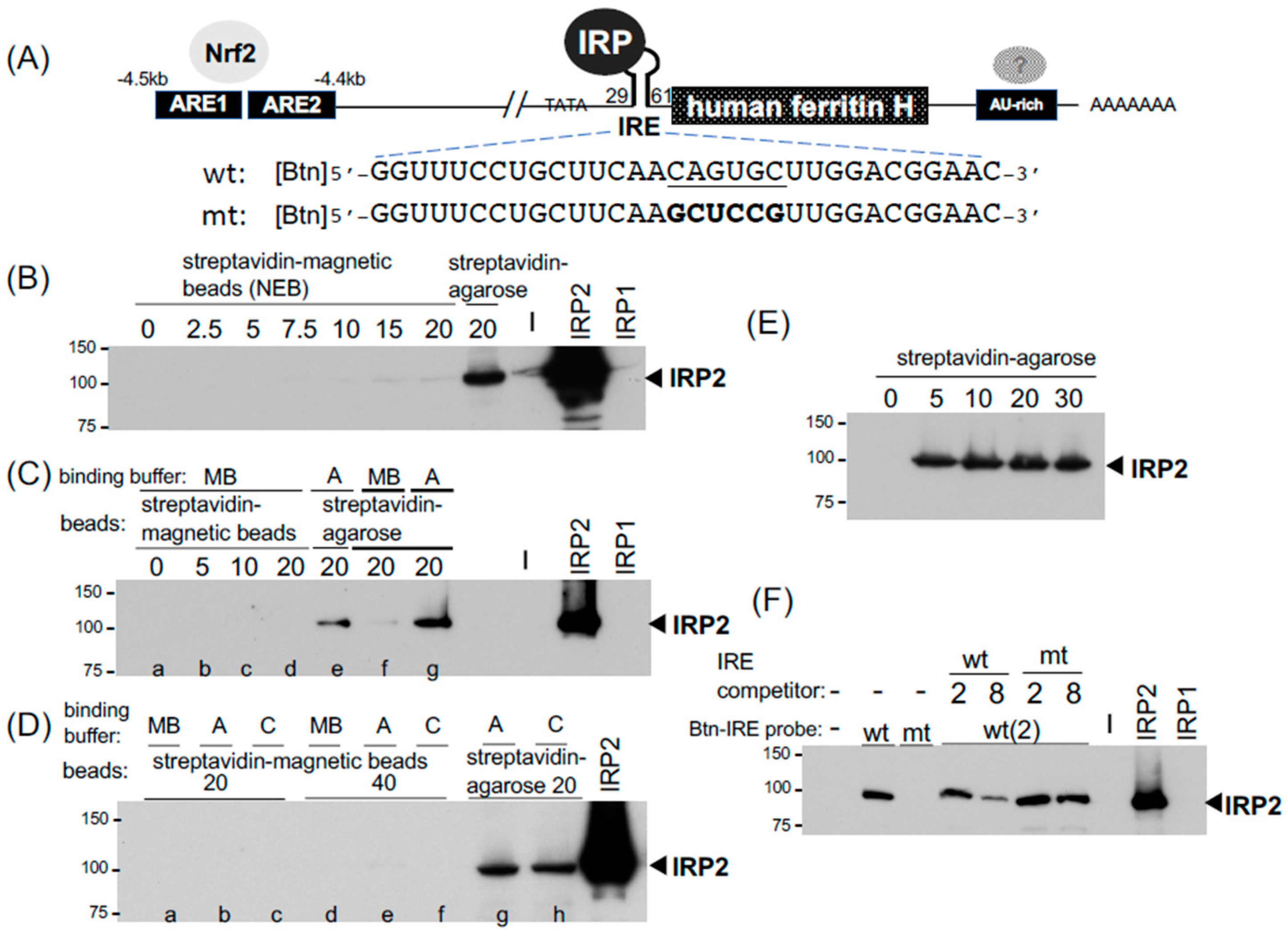
IJMS | Free Full-Text | Optimization of Biotinylated RNA or DNA Pull-Down Assays for Detection of Binding Proteins: Examples of IRP1, IRP2, HuR, AUF1, and Nrf2
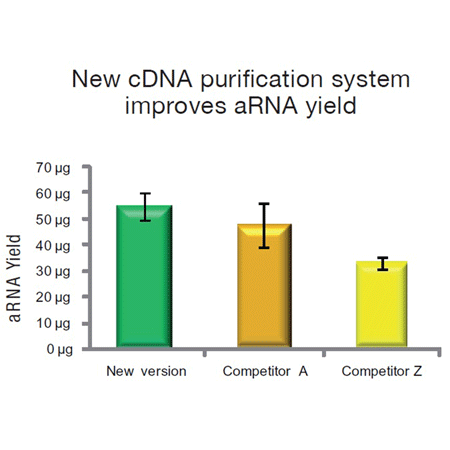
BIOARRAY™ Single-round RNA amplification and biotin labeling system - ENZ-42420 - Enzo Life Sciences

EU-RNA-seq for in vivo labeling and high throughput sequencing of nascent transcripts - ScienceDirect

Synthesis of biotin–AMP conjugate for 5′ biotin labeling of RNA through one-step in vitro transcription | Nature Protocols
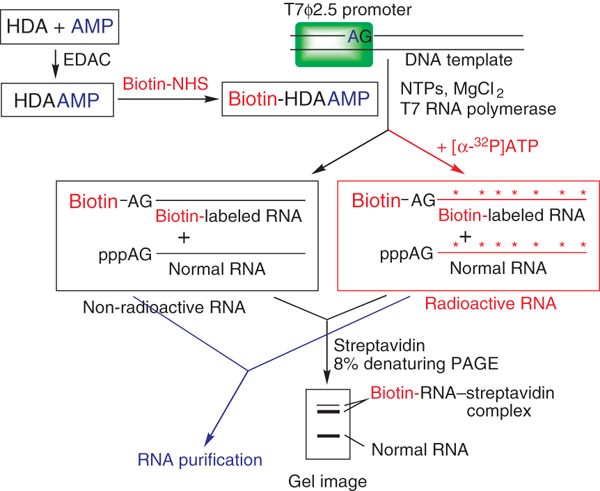
Synthesis of biotin–AMP conjugate for 5′ biotin labeling of RNA through one-step in vitro transcription | Nature Protocols

Hybridization-proximity labeling reveals spatially ordered interactions of nuclear RNA compartments - ScienceDirect

Enzymatic RNA Biotinylation for Affinity Purification and Identification of RNA-protein Interactions | bioRxiv
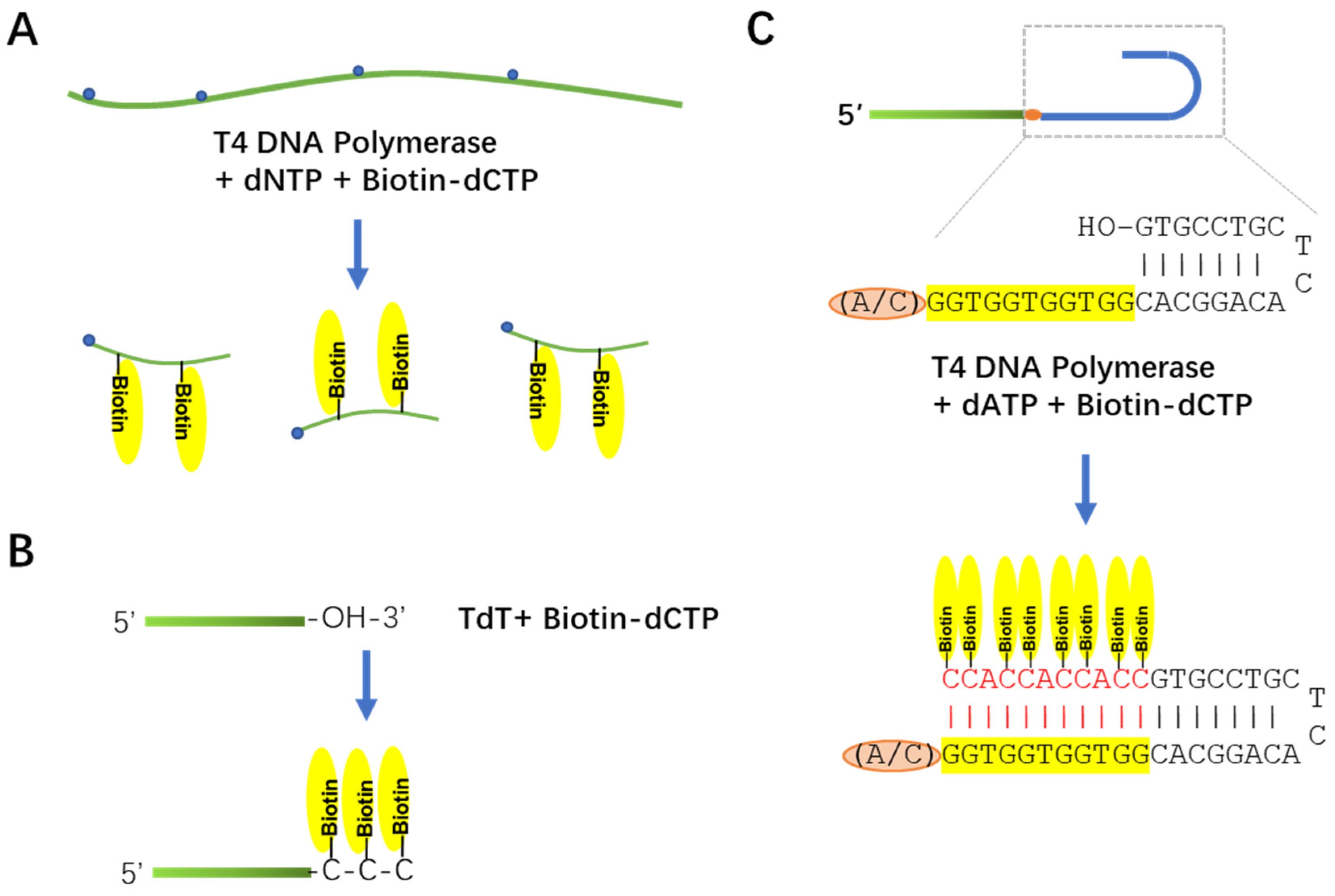
Viruses | Free Full-Text | A New Biotin Labeling and High-Molecular-Weight RNA Northern Method and Its Application in Viral RNA Detection
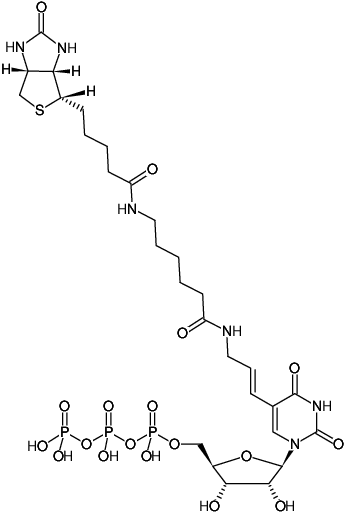

HighYield T7 Biotin11 RNA Labeling Kit (UTP-based), Random RNA Labeling (in vitro Transcription-based) - Jena Bioscience